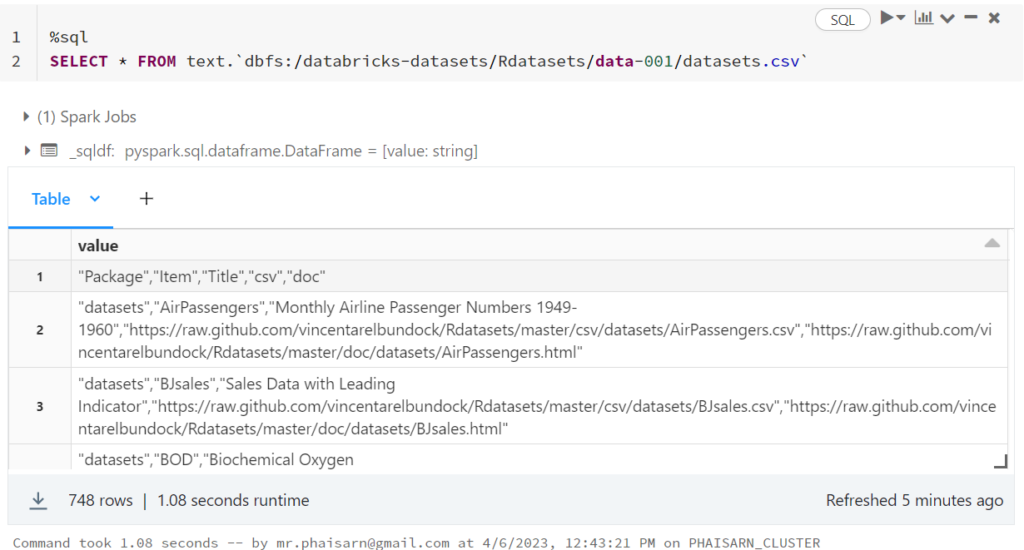
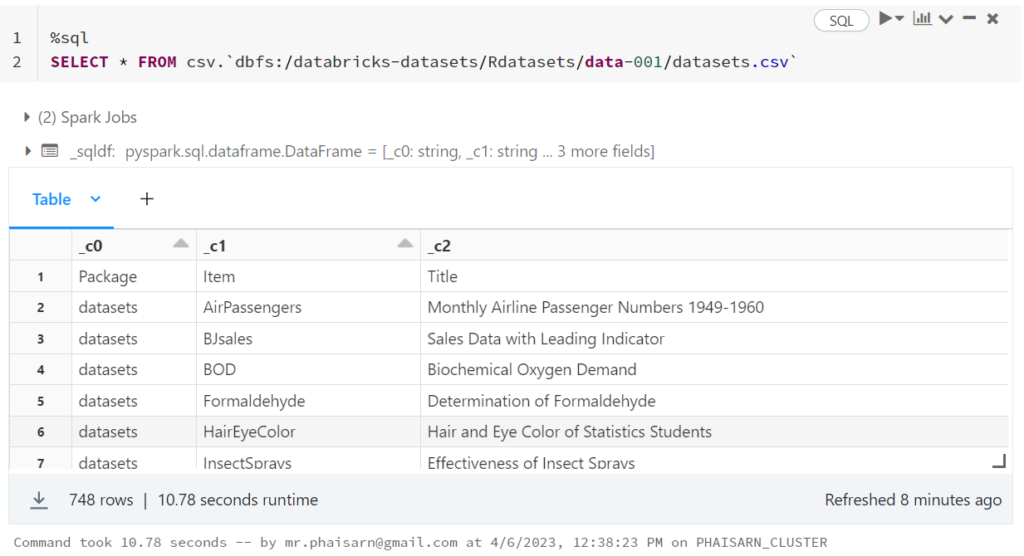
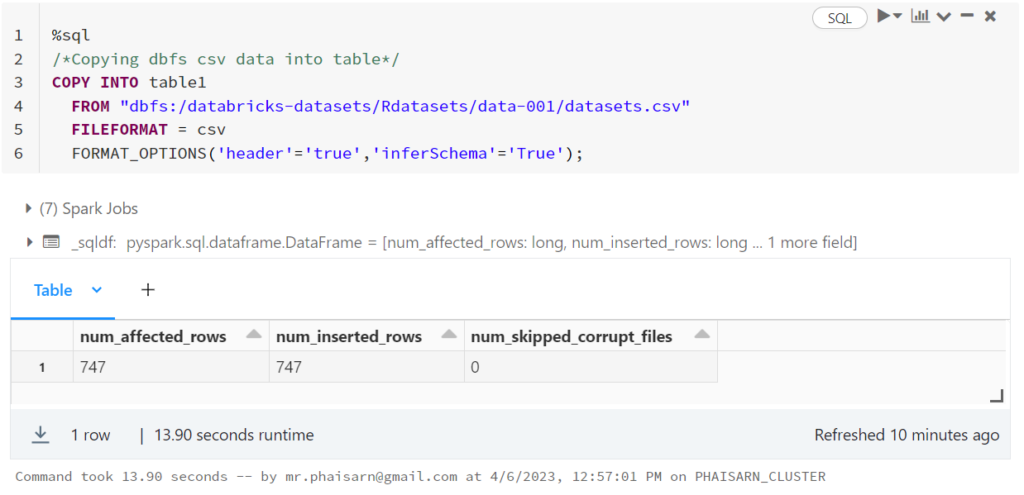
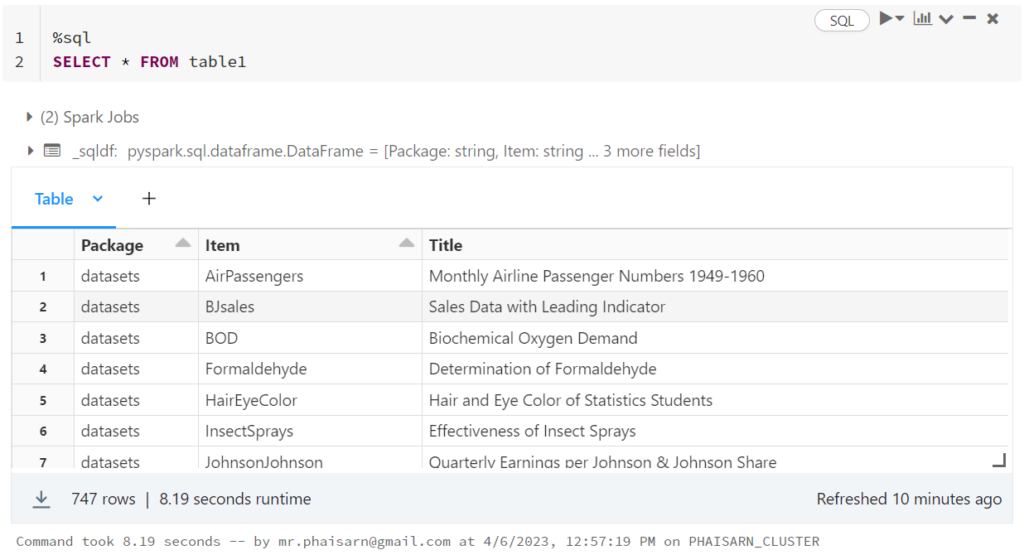
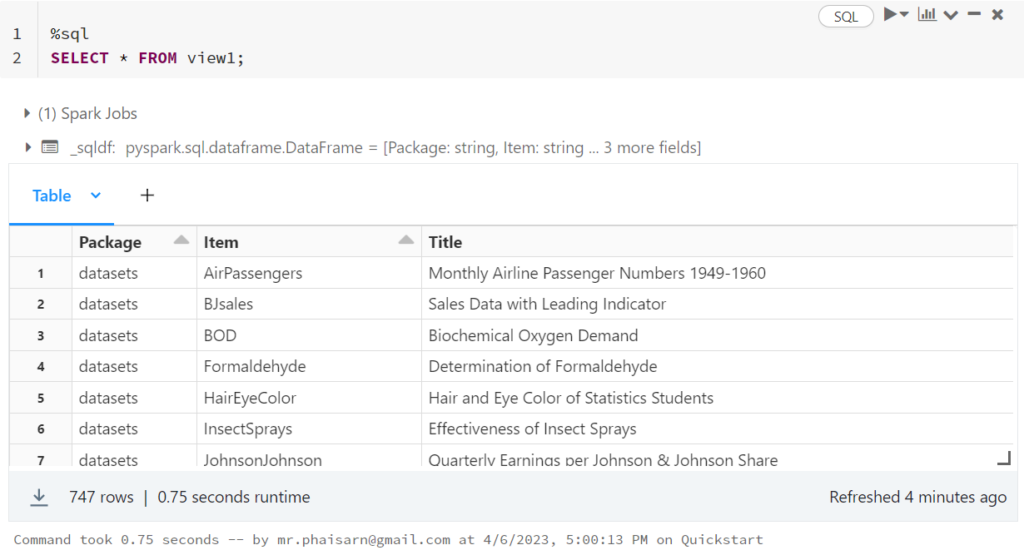
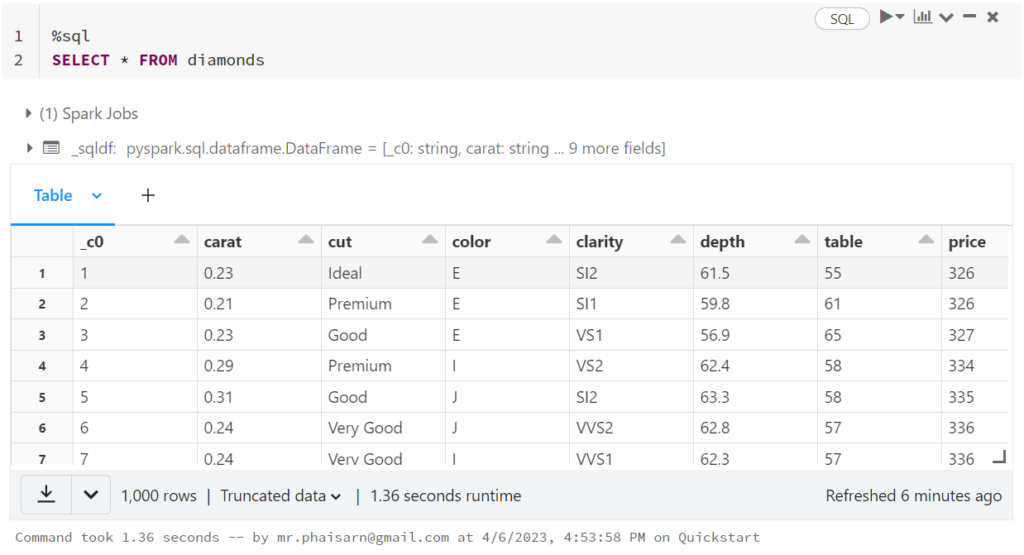
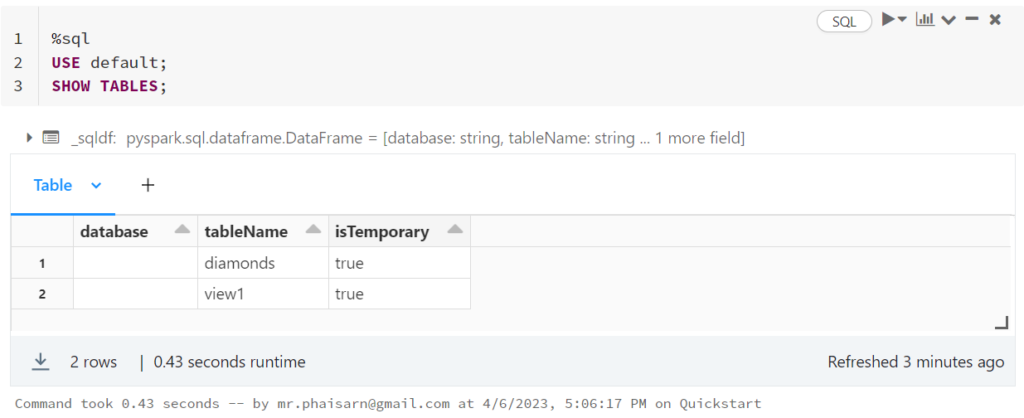
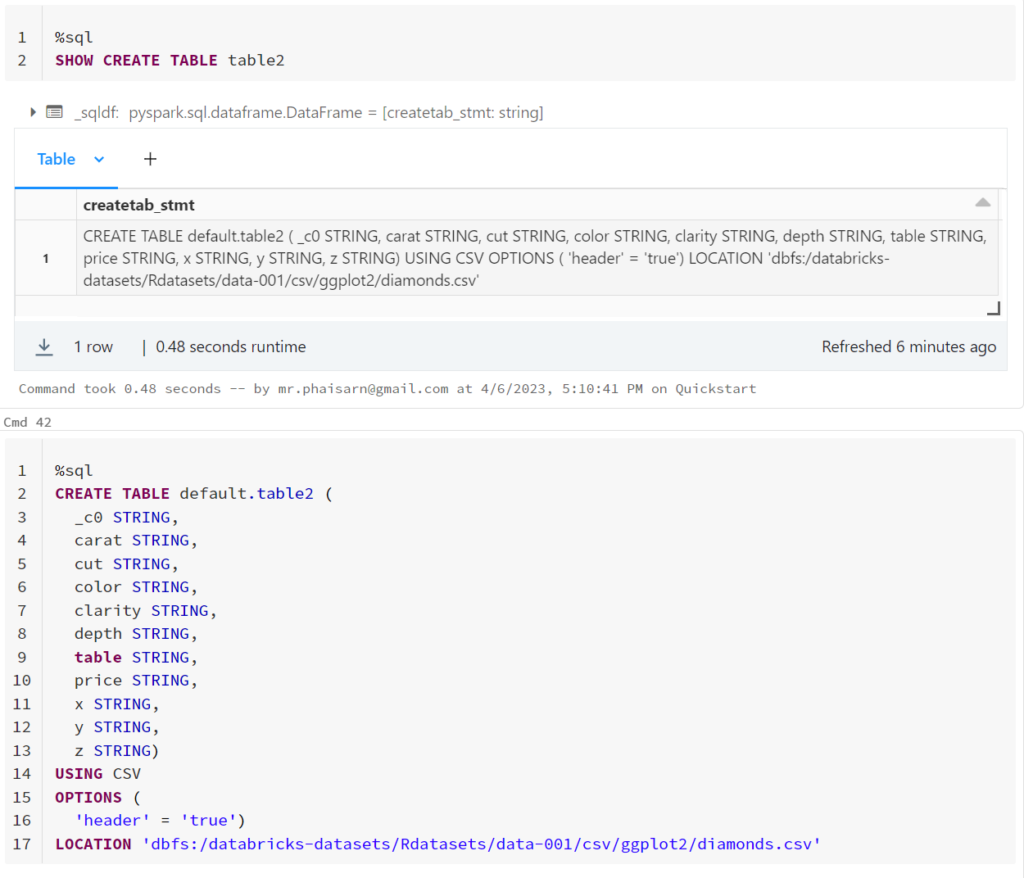
1. สร้างตาราง students
%sql CREATE TABLE students ( id INT, name STRING, value DOUBLE);
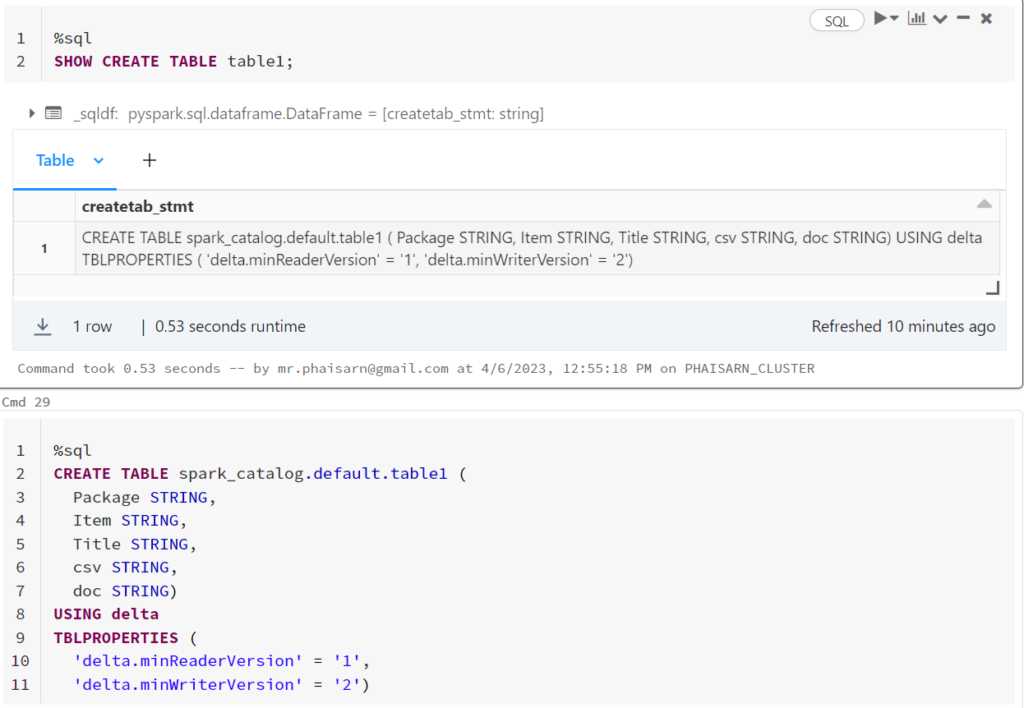
ดูคำสั่งสร้างตารางนี้ด้วย SHOW CREATE TABLE
%sql SHOW CREATE TABLE students
%sql CREATE TABLE spark_catalog.default.students ( id INT, name STRING, value DOUBLE) USING delta TBLPROPERTIES ( 'delta.minReaderVersion' = '1', 'delta.minWriterVersion' = '2')
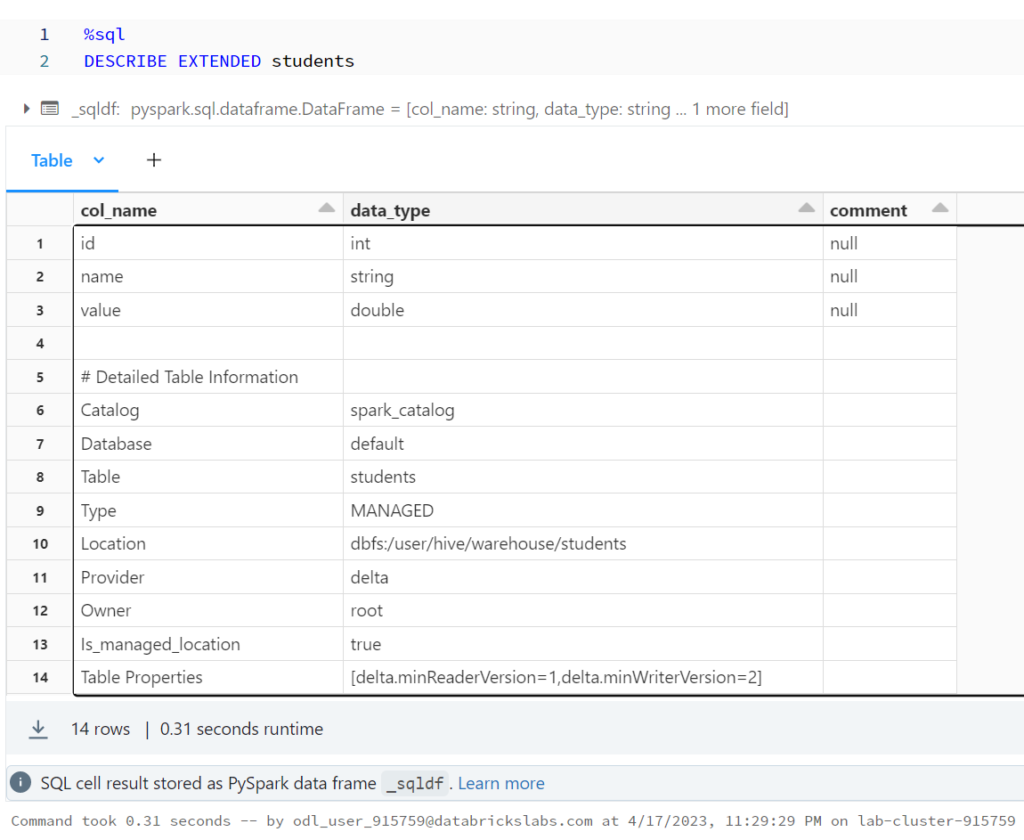
Using DESCRIBE EXTENDED allows us to see important metadata about our table.
%sql DESCRIBE EXTENDED students
%python
df1 = spark.sql('DESCRIBE EXTENDED students')
df1.show()
+--------------------+--------------------+-------+ | col_name| data_type|comment| +--------------------+--------------------+-------+ | id| int| null| | name| string| null| | value| double| null| | | | | |# Detailed Table ...| | | | Catalog| spark_catalog| | | Database| default| | | Table| students| | | Type| MANAGED| | | Location|dbfs:/user/hive/w...| | | Provider| delta| | | Owner| root| | | Is_managed_location| true| | | Table Properties|[delta.minReaderV...| | +--------------------+--------------------+-------+
DESCRIBE DETAIL is another command that allows us to explore table metadata.
%sql DESCRIBE DETAIL students
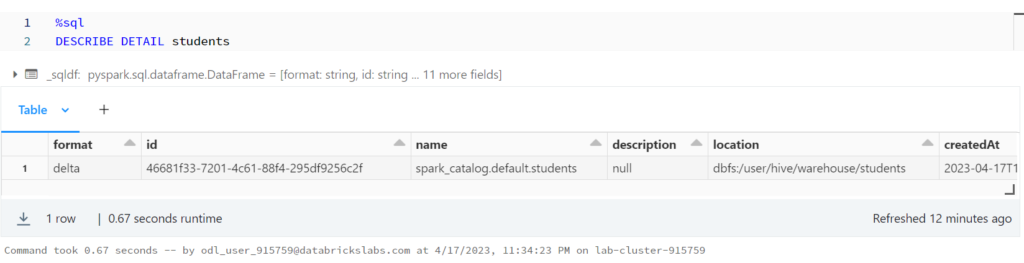
%python
df2 = spark.sql('DESCRIBE DETAIL students')
df2.show(vertical=True)
-RECORD 0--------------------------------
format | delta
id | 46681f33-7201-4c6...
name | spark_catalog.def...
description | null
location | dbfs:/user/hive/w...
createdAt | 2023-04-17 16:24:...
lastModified | 2023-04-17 16:24:32
partitionColumns | []
numFiles | 0
sizeInBytes | 0
properties | {}
minReaderVersion | 1
minWriterVersion | 2
ดู Delta Lake Files
%python
display(dbutils.fs.ls('dbfs:/user/hive/warehouse/students'))
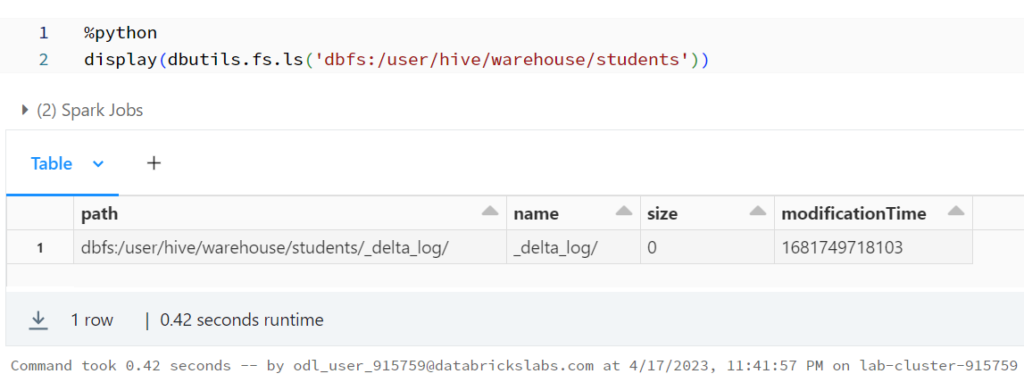
%python
li_file = dbutils.fs.ls('dbfs:/user/hive/warehouse/students')
df3 = sqlContext.createDataFrame(li_file)
df3.show()
+--------------------+--------------------+----+----------------+ | path| name|size|modificationTime| +--------------------+--------------------+----+----------------+ |dbfs:/user/hive/w...| _delta_log/| 0| 1681750558289| +--------------------+--------------------+----+----------------+
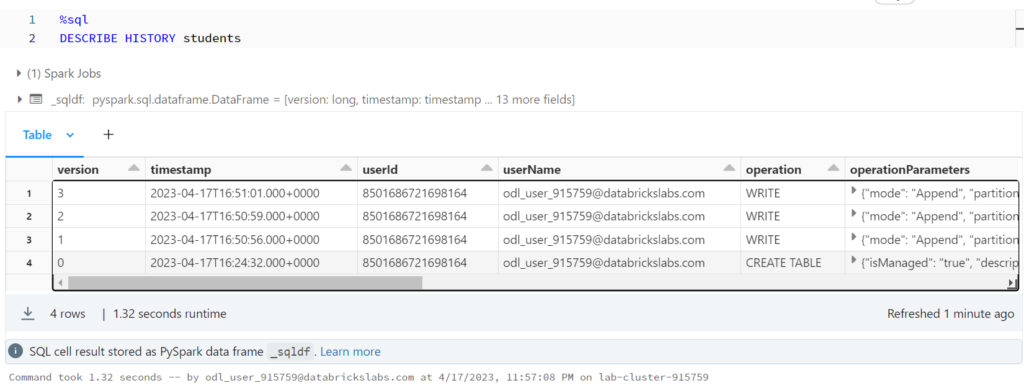
Reviewing Delta Lake Transactions
%sql DESCRIBE HISTORY students
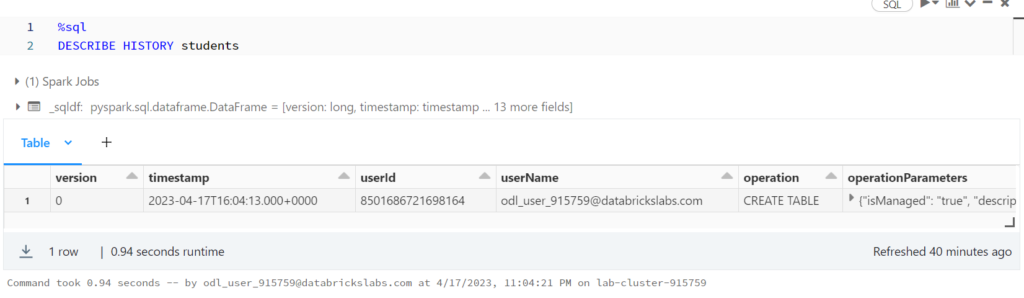
%python
df4 = spark.sql('DESCRIBE HISTORY students')
df4.show(vertical=True)
-RECORD 0-----------------------------------
version | 0
timestamp | 2023-04-17 16:24:32
userId | 8501686721698164
userName | odl_user_915759@d...
operation | CREATE TABLE
operationParameters | {isManaged -> tru...
job | null
notebook | {1477724271071511}
clusterId | 0415-162149-6ai590aw
readVersion | null
isolationLevel | WriteSerializable
isBlindAppend | true
operationMetrics | {}
userMetadata | null
engineInfo | Databricks-Runtim...
2. เพิ่มข้อมูล 3 ครั้ง
%sql INSERT INTO students VALUES (1, "Yve", 1.0); INSERT INTO students VALUES (2, "Omar", 2.5); INSERT INTO students VALUES (3, "Elia", 3.3);
%python
df2 = spark.sql('DESCRIBE DETAIL students')
df2.show(vertical=True)
RECORD 0--------------------------------
format | delta
id | 46681f33-7201-4c6...
name | spark_catalog.def...
description | null
location | dbfs:/user/hive/w...
createdAt | 2023-04-17 16:24:...
lastModified | 2023-04-17 16:51:01
partitionColumns | []
numFiles | 3
sizeInBytes | 2613
properties | {}
minReaderVersion | 1
minWriterVersion | 2
%python
li_file = dbutils.fs.ls('dbfs:/user/hive/warehouse/students')
df3 = sqlContext.createDataFrame(li_file)
df3.show()
+--------------------+--------------------+----+----------------+ | path| name|size|modificationTime| +--------------------+--------------------+----+----------------+ |dbfs:/user/hive/w...| _delta_log/| 0| 1681750558289| |dbfs:/user/hive/w...|part-00000-1d6df3...| 868| 1681750256000| |dbfs:/user/hive/w...|part-00000-57bc12...| 872| 1681750261000| |dbfs:/user/hive/w...|part-00000-ec8db3...| 873| 1681750259000| +--------------------+--------------------+----+----------------+
%python display(spark.sql(f"SELECT * FROM json.`dbfs:/user/hive/warehouse/students/_delta_log/00000000000000000001.json`"))
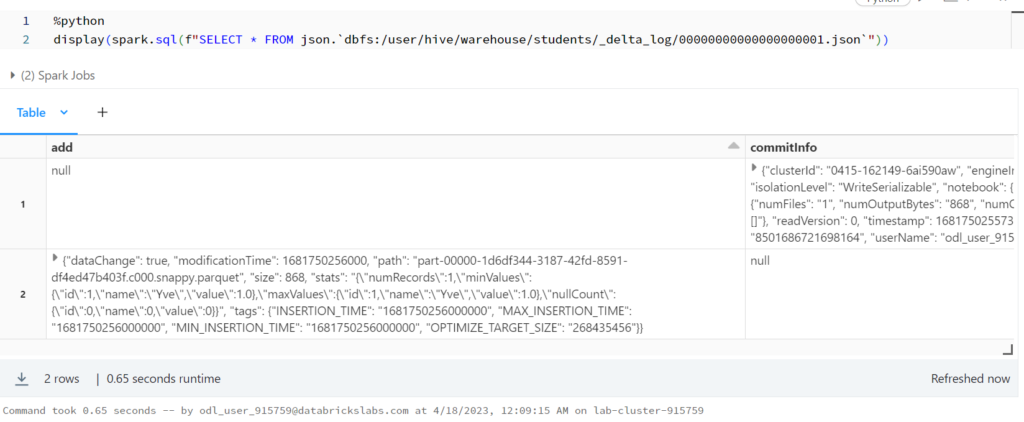
รันคำสั่งนี้จะ error
%sql SELECT * FROM parquet.`dbfs:/user/hive/warehouse/students/part-00000-1d6df344-3187-42fd-8591-df4ed47b403f.c000.snappy.parquet`
AnalysisException: Incompatible format detected.
A transaction log for Delta was found at `dbfs:/user/hive/warehouse/students/_delta_log`,
but you are trying to read from `dbfs:/user/hive/warehouse/students/part-00000-57bc1277-813b-41c0-990d-4ba8fd05c9b1.c000.snappy.parquet` using format("parquet"). You must use
'format("delta")' when reading and writing to a delta table.
To disable this check, SET spark.databricks.delta.formatCheck.enabled=false
To learn more about Delta, see https://docs.databricks.com/delta/index.html; line 1 pos 14
ให้ SET ค่านี้ก่อน
%sql SET spark.databricks.delta.formatCheck.enabled=false
รันใหม่จะได้ละ
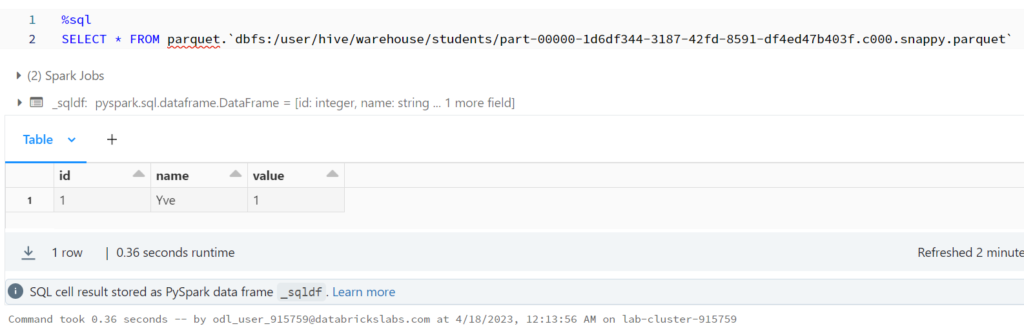
%sql SELECT * FROM parquet.`dbfs:/user/hive/warehouse/students/part-00000-1d6df344-3187-42fd-8591-df4ed47b403f.c000.snappy.parquet`